We are thrilled to announce that our latest paper, “ProFiT-SPEci-FISH,” has been accepted for publication in Microbiome!
Plasmids are small, circular DNA molecules that are the heavy lifters of bacterial evolution. They carry critical traits like antibiotic resistance and other metabolic capabilities, jumping between bacteria via horizontal gene transfer (HGT). However, studying natural plasmids has always presented a paradox:
Metagenomic sequencing allows us to see which plasmids are present, but we often lose the context of their bacterial hosts.
In our new paper, we introduce a pipeline that bridges this gap, allowing us to link plasmids to their specific bacterial hosts in complex, natural environments at the single-cell level. It combines computational design with a robust wet-lab technique:
SPEci-FISH (Short Probe EffiCIent Fluorescence In Situ Hybridization): This molecular method uses the probes from ProFiT combined with enzymatic signal amplification. This allows us to visualize plasmids even at low copy numbers with exceptional sensitivity and specificity.
ProFiT (PRObe FInding Tool): This is our new bioinformatic tool. It scans the metagenome of the specific sample to design multiple short, specific probes. It ensures these probes target only the plasmid of interest, minimizing false positives.

This combined pipeline is versatile, allowing for both microscopy-based visualization and high-throughput screening via fluorescence-activated cell sorting (FACS). We tested this pipeline on both clinical isolates and the complex environment of the human gut.
What did we find?
1. Inside the Cell: Distinct Spatial Patterns
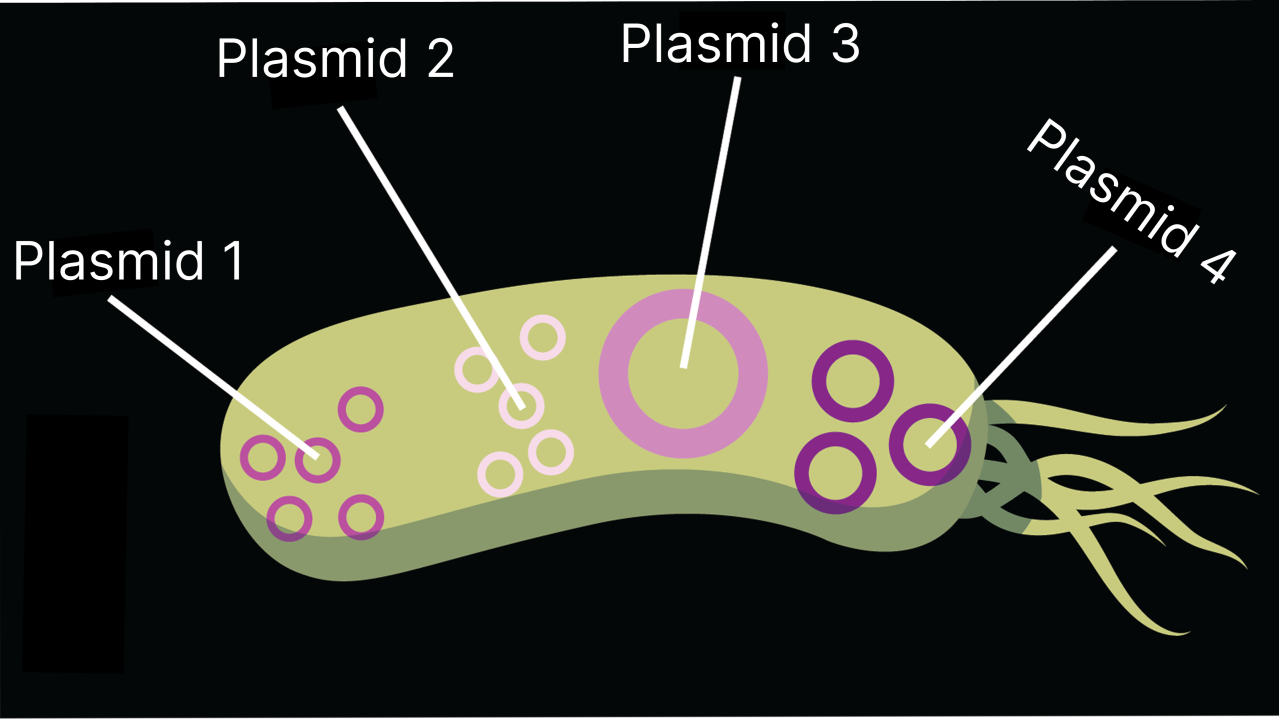
Using super-resolution microscopy (STORM), we visualized four different plasmids within a multidrug-resistant E. coli clinical isolate. We observed something fascinating: while some plasmids distributed randomly, one specific plasmid clustered at the cell poles. This suggests that different plasmids within the same cell utilize different segregation mechanisms.
2. In the Wild: Identifying Hosts in the Human Gut
The true test came when we applied the pipeline to a human fecal sample. Using bioinformatics, we identified a conjugative plasmid, a genetic element capable of transferring itself between bacteria to spread new traits, and used ProFiT-SPEci-FISH to track it. By combining our labeling method with Fluorescence-Activated Cell Sorting (FACS), we isolated and sequenced the specific cells carrying the plasmid. We discovered that the plasmid acts as a promiscuous traveler, residing in two phylogenetically distant hosts: Bacteroides fragilis and Ruminococcus bromii.
We found that while the plasmid was present in both, it had a higher prevalence in B. fragilis compared to R. bromii, suggesting a source-sink dynamic where B. fragilis acts as the primary reservoir.

These images show specific DNA circles (called pE2022 plasmids) inside individual bacteria. The colors represent how crowded the DNA is in different areas: white shows where the DNA is most tightly packed, while black shows where there is very little. As the arrows point out, these DNA circles tend to bunch up at the very ends of the cells.
Why This Matters?
ProFiT-SPEci-FISH allows us to move beyond metagenomic sequencing. We can now identify the exact bacterial hosts of plasmids in their native habitats without needing to culture them first. This is a significant step forward in understanding how antibiotic resistance spreads and how microbial communities evolve.
The ProFiT tool is open-source and available on GitHub for anyone interested in exploring plasmid-host dynamics in their own samples! – https://github.com/labmizrahi/ProFiT
Read the full story here: https://link.springer.com/article/10.1186/s40168-025-02238-z
to the Benyamin Rosental & Natalie Elia Group for an amazing collaboration!
A huge congratulations to our lead author, Alvah Zorea who spearheaded this study. Huge thanks to the amazing contributors: Sarah Morais, David Pellow, Orly Gershoni-Yahalom, Maraike Probst, Sapir Nadler & Ron Shamir