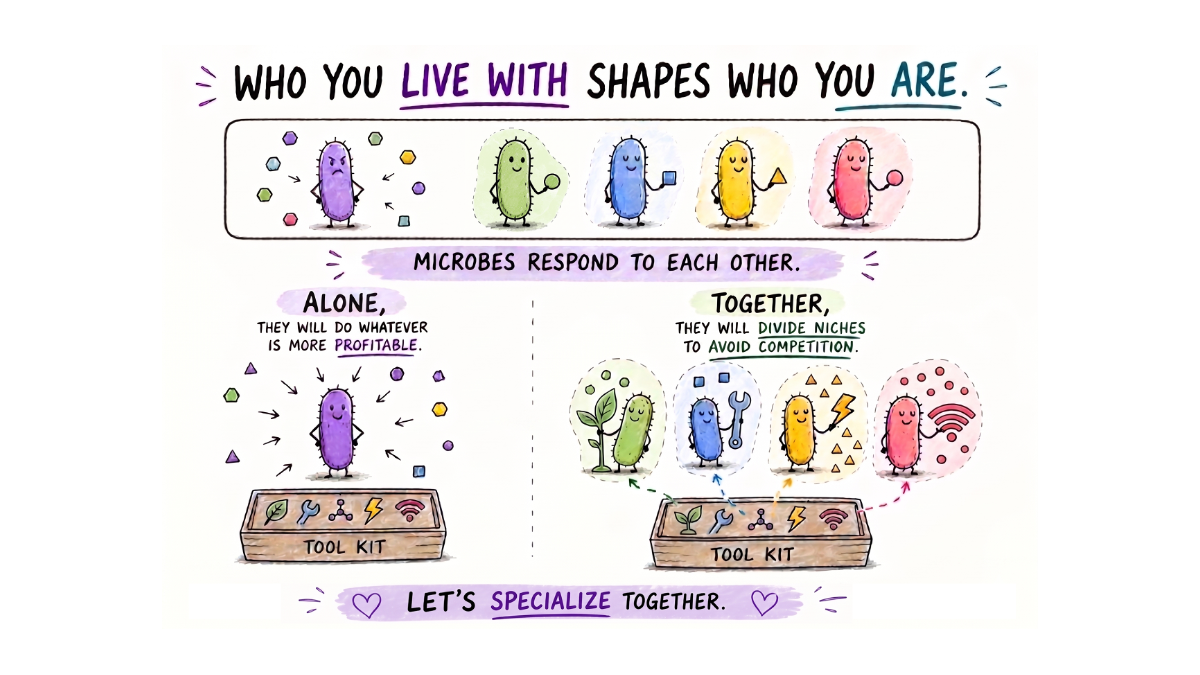
Do microbes behave differently when they are not alone?
Most of us think about microbes as individual species. This bacterium does one thing, that bacterium does another.
But in nature, microbes are almost never alone.
They live in dense, crowded communities, in our gut, in the rumen of cows, in soil, in oceans, essentially everywhere. And like people sharing a workspace, they need to figure out how to coexist without everyone doing the same job, wasting energy, and competing for the same resources.
In our new paper, published in Nature Microbiology, we asked a simple but fundamental question:
Do microbes change what they do depending on who is around them?
The answer is yes, and the changes are striking.
We built small, controlled microbial communities using gut bacteria, and instead of simply asking which microbes were present, we examined which proteins each microbe was producing. Proteins are the active molecular tools that cells use to carry out specific tasks, so this gave us a direct, real-time view of what each bacterium was actually doing.
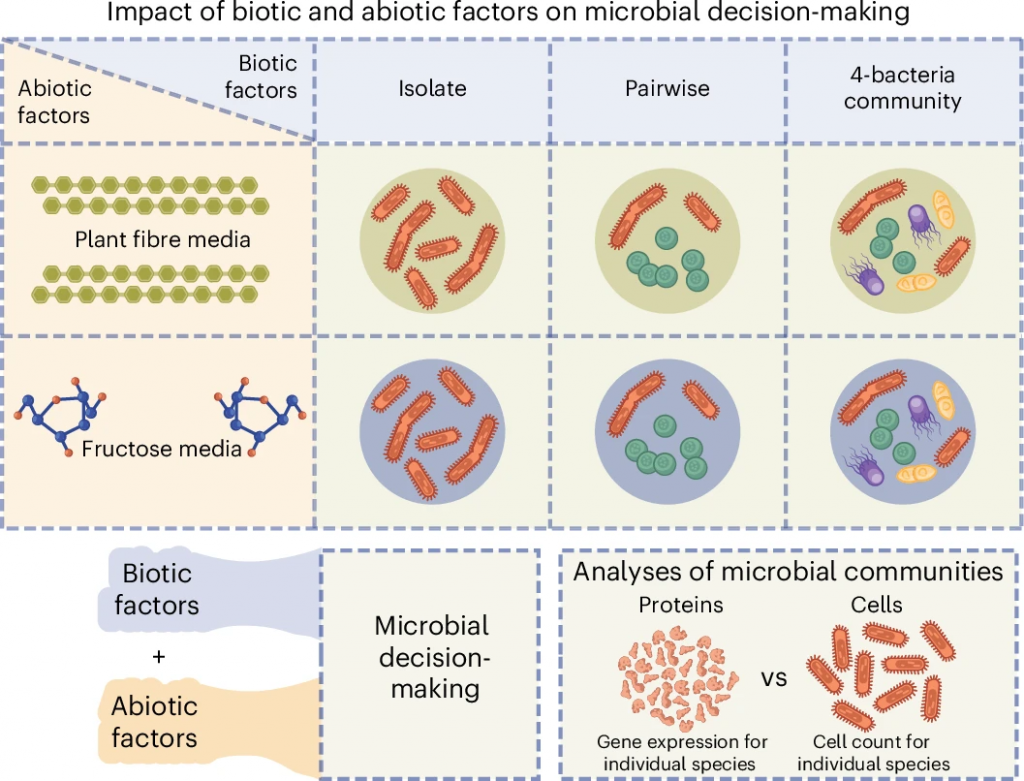
What we found was surprising….
The identity of neighboring microbes had a stronger effect on protein production than the food source itself. In other words, microbes were not only responding to what they were eating, they were responding even stronger to who they were living with.
Our in-depth protein expression analyses allowed us to uncover a potential underlying mechanism: when microbes lived together, they reduced functional overlap. Instead of all doing the same things, they appeared to specialize. Core essential functions were maintained, but many overlapping activities were reduced.
It’s a bit like an ensemble, where not everyone brings the same tools or plays the same role. Instead, each member contributes something different, avoiding unnecessary competition and becoming more efficient together.
That’s exactly what we observed. This reduction in overlap was often associated with higher community productivity.
Of course, this happens at the molecular level, most likely through sensing and responding to the metabolites produced by their neighbors.
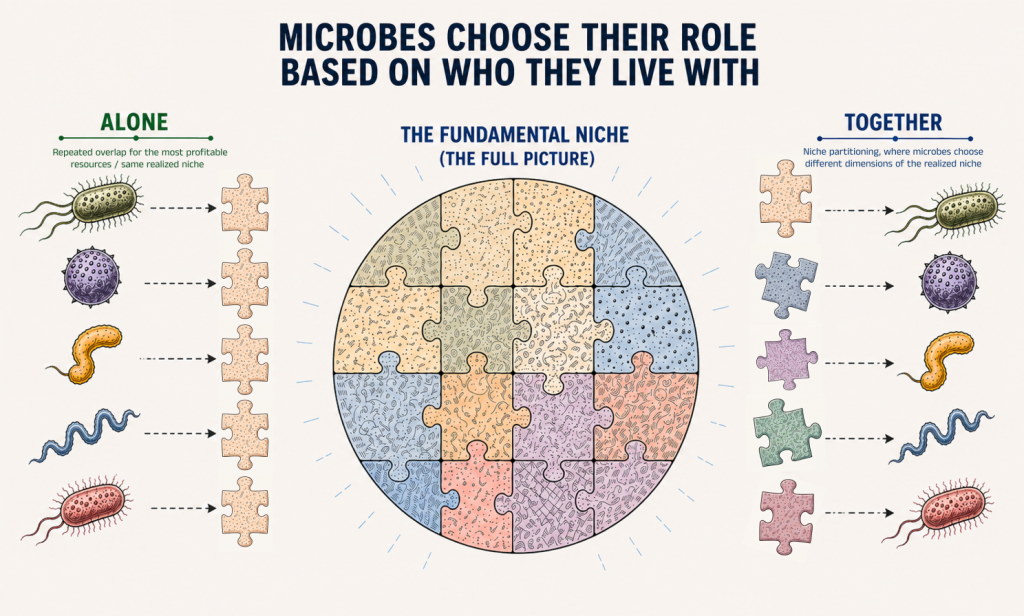
To me, this is particularly exciting because it provides a molecular explanation for a classic ecological idea: how organisms live together in communities. Instead of competing for the same resources, they coexist by dividing them, reducing competition, and enabling more efficient use of what is available.
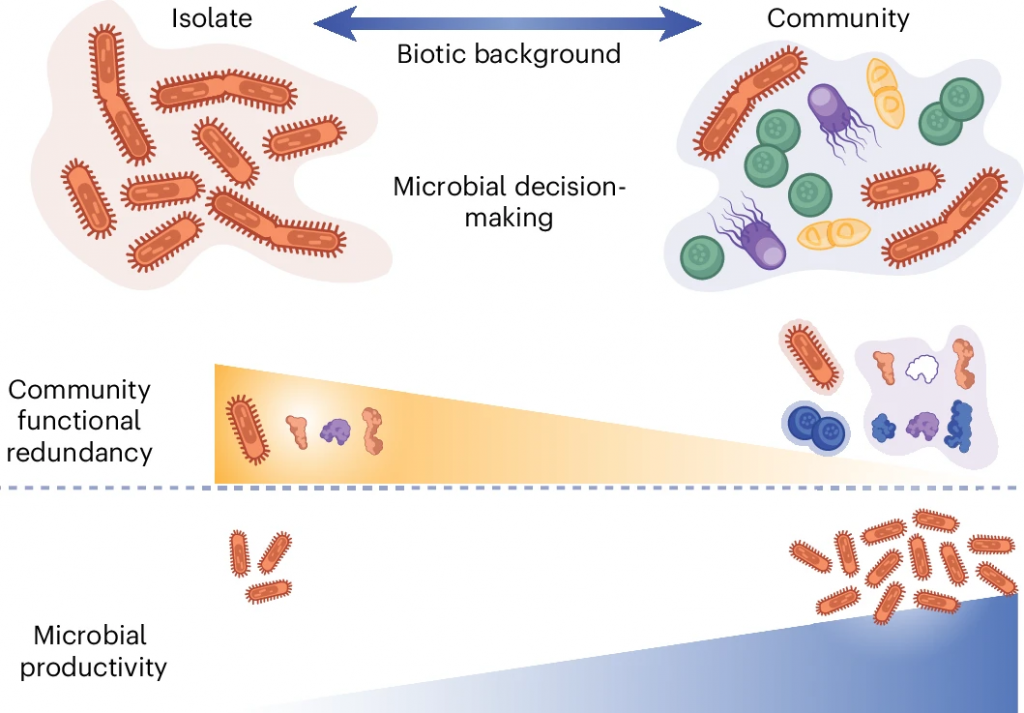
Microbes are not just passively coexisting.
They are actively responding and adjusting to each other.
This means that a microbe is not defined only by its genome, which represents its potential, but also by its community.
The same bacterium can behave very differently depending on who surrounds it.

Why this matters beyond basic science
If microbes actively coordinate what they do in response to each other, then microbiomes are not just collections of species. They are dynamic, adaptive systems.
And that shift in perspective matters.
In human health, it suggests that designing probiotics or microbiome-based therapies is not just about choosing the “right” microbes. It is about finding the right combinations, communities where microbes naturally divide tasks instead of competing or duplicating effort.
In agriculture, especially in systems like the rumen where microbes control feed efficiency and methane emissions, understanding how communities organize themselves could help us guide them toward more productive and sustainable states.
In biotechnology, it points toward a different strategy. Instead of engineering single “super microbes,” we can design ensembles of microbes, each doing part of the job, making the whole system more efficient and stable.
And in microbiome restoration, it offers a new explanation for why some microbes fail to establish. It may not be that they are missing, but that their ecological “home” is missing. Rebuilding the right community context could allow them to return.
More broadly, our work suggests that microbial communities follow underlying rules. Division of labor and efficiency emerge naturally from interactions and is manifasted at the protein expression level.
If we can understand these rules, we move from simply describing microbiomes to actually predicting, shaping, and engineering them.
This work was led by Sarah Moraïs together with an amazing team across Ben-Gurion University and the University of Greifswald, Michael Mazor , itai amit , Ido Grinshpan, Philip Gerth, Anke Trautwein-Schult, , Sandra Maaß, Yehonatan Shelly, Liron Levin, and Dörte Becher, you guys are awesome.
The research was supported by the European Research Council (ERC 866530), the Israel Science Foundation–Swiss National Science Foundation (ISF–SNSF 1057/24), and the Israel Science Foundation (979/25).
Very proud to share this paper:
Community context reshapes microbial proteomes and reduces functional overlap
Published in Nature Microbiology